Protocol
Goal :
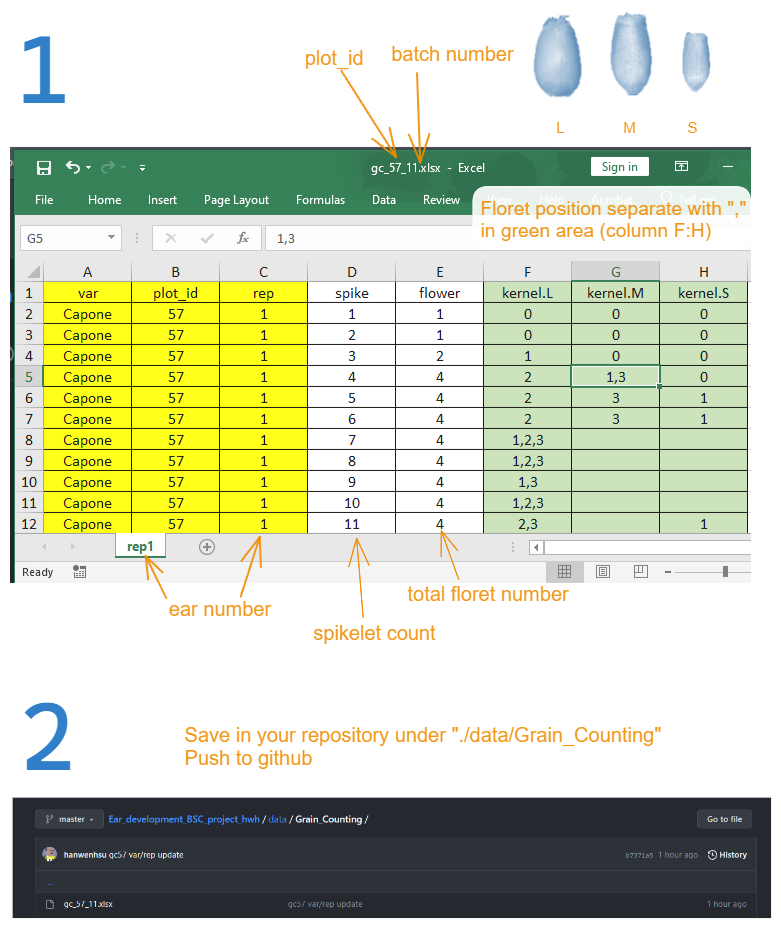
- Treatment to compare: early and late
- Other info (total nitrogen 176 kg/ha, split application of heading fertilizer, replicate 1).
- Cultivar: Student A(Capone) Student B (Patras, Pionier) Student C (Potenzial)
- Date: 5 batches from 11.

Protocol
- Clean up the table. Place one A4 paper on the table.
- Take out one ear at a time from a bag containing 10 ears.
- download grain_counting_example.xlsx from HuBox
- Copy to your repository under relative path
./data/Grain_Counting/save it asgc_plotid_batch.xlsx(Figure 2) - Enter information of
var,plot_idandear number(from 1 to 10) - Count total
spikelet number (Nsp), record the sequence 1:Nspin columnspike. - Count the
floret number (Nf)based onNspwith ascending order (start from spikelet 1). - Classify the shape of fully developed kernel into three classes (
L,M,S), record the position of each class, separate them with comma (,). For example: 1,2,3 for classkernel.L(Figure 2) - Aborted kernel will not be recorded in column
F:H. However, if special condition observed, you can record it in columnI(with column namenote) and take a picture of it. - Roll your ear with another half-A4 paper like a candy. Make sure you have the ear number and plotid on the outer side of the paper. And put it back to the bag.
- Check the correctness and completeness of the data in current sheet. Save the data, start another ear in a new sheet.
- Get up and take a small break!

Challenge: automatize process of reading files
Use the code below to reach the following goals.
- Write a
for loopto read all the files and sheets in the folder - add batch information to column
library(dplyr)
p <- ".data/Grain_Counting/gc_57_11.xlsx"
df <- readxl::read_xlsx(p) %>%
mutate(across(starts_with("kernel"),function(x)as.character(x))) %>%
tidyr::pivot_longer(starts_with("kernel"),names_to = "kernel.type",values_to = "floret.pos") %>%
mutate(floret.pos=strsplit(floret.pos,",")) %>%
tidyr::unnest(floret.pos) %>%
mutate(floret.pos=as.numeric(floret.pos) %>%replace(., is.na(.), 0))